New publication, Imane Doghman, et al. in J. Med. Virol.
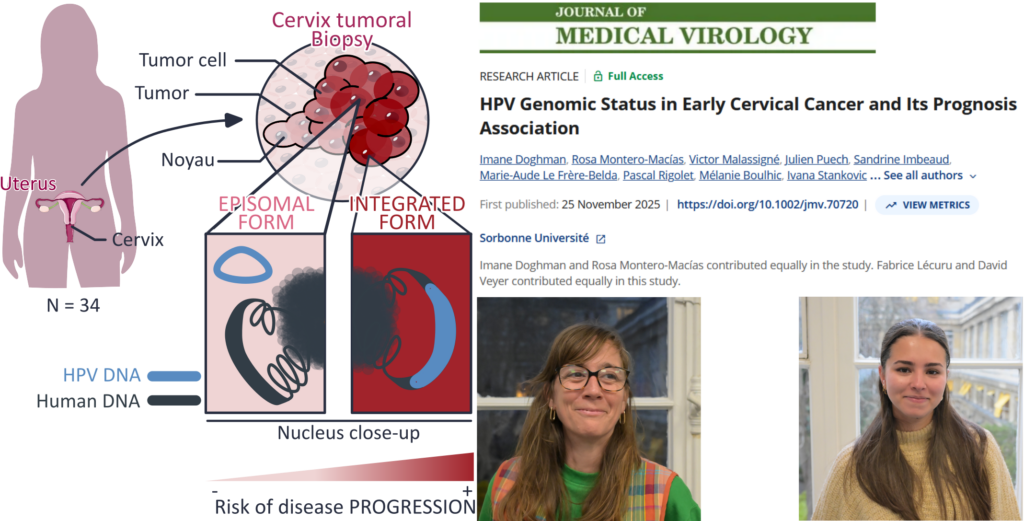
Identify specific Human Papilloma Virus (HPV) genomic status by HPV capture technique coupled with next-generation sequencing (NGS) in patients with early cervical cancer (ECC) regarding 2018 FIGO stages and correlate these statuses with prognosis. HPV sequencing by Capture technique coupled with NGS was performed on 34 tumors samples of ECC patients without histologically sentinel lymph node (SLN) metastasis in a retrospective study. NGS data were analyzed in our in-house VIROCAPT2 pipeline to determine precise HPV genomic status such as the complete sequence of HPV genome, its integration or not in host genome and correlate this viral identity card with patient’s clinical prognosis. As expected, HPV16 remains the most common genotype (60%, n = 21/35) in ECC. HPV integration was identified in 40% (n = 14/35) of samples. We found that in ECC, integration events were related with HPV genotype (p = 0.0005). Among the 40% of integrated HPV, 8 were found integrated within human genes, including 2 into oncogenes. Finally, women with integrated HPV had a poorest outcome with a strong correlation between progression-free survival and HPV integration (p = 0.0035) and interestingly, the 2 HPV integrated into oncogenes were observed in patients who died from cancer. HPV genomic status seems to be a key prognostic factor in early stage cervical cancer, with HPV integration strongly linked to poor outcomes. Larger studies are needed to confirm the clinical relevance of HPV ID card for risk stratification and treatment decisions.